FLARE: Measurement System
Introduction
We propose FLARE for plant disease severity assessment, consisting of FLLA-Net and ERMA-Net. FLLA-Net uses a dual-stream segmentation network to extract leaf and lesion features, while ERMA-Net leverages these features and applies relational modeling to capture spatial and semantic relationships. Enhanced with global feature refinement and contrastive learning, the framework predicts severity levels accurately, providing a robust solution under complex field conditions.
Dependencies
- CUDA 11.8
- Python 3.8 (or later)
- torch==1.13.1
- torchaudio==0.13.1
- torchcam==0.3.2
- torchgeo==0.4.1
- torchmetrics==0.11.4
- torchvision==0.14.1
- numpy==1.21.6
- Pillow==9.2.0
- einops==0.6.0
- opencv-python==4.6.0.66
GitHub: https://github.com/GZU-SAMLab/FLARE
💾 Code and Data
The source code can be downloaded from this link.
Only the test set of the plant leaf disease severity assessment dataset is publicly available for now. The training set will be released once the paper is accepted. You can download the dataset from this link. Download the dataset to the './dataset/' folder.
🧩 Pre-trained Models
The pre-trained FLLA-Net_Leaf, FLLA-Net_Lesion, and ERMA-Net models are provided below.
Please download them and place the files into the
./checkpoint folder.
You can download all pre-trained weights from
this link.
Get Started
🧠 Train DLAR_Net
Step 1:Train FLLA-Net_Leaf
python trainUNet++.py
Step 2:Train FLLA-Net_Lesion
python trainDLAR_Net.py
⚙️ Train ERMA-Net
python trainL2RA_Net.py --epochs 300 --batch_size 16 \
--model_path_lesion ./checkpoint/FLLA-Net_Lesion.pth \
--model_path_leaf ./checkpoint/FLLA-Net_Leaf.pth
🚀 Inference
python trainL2RA_Val.py --model_path_classifier ./checkpoint/ERMA-Net.pth
Results
Quantitative Analysis
To validate the proposed FLARE framework, FLLA-Net is first pre-trained on a leaf–lesion segmentation dataset, followed by training ERMA-Net on the severity dataset with corresponding image descriptions. As shown in Figure 1, FLARE achieves the highest overall accuracy, outperforming 15 existing models by at least 3.11%. Its ability to explicitly model lesion–leaf relationships enables precise and human-consistent severity estimation. In contrast, ViT shows strong bias toward the first severity level, indicating limited discrimination across multiple grades.
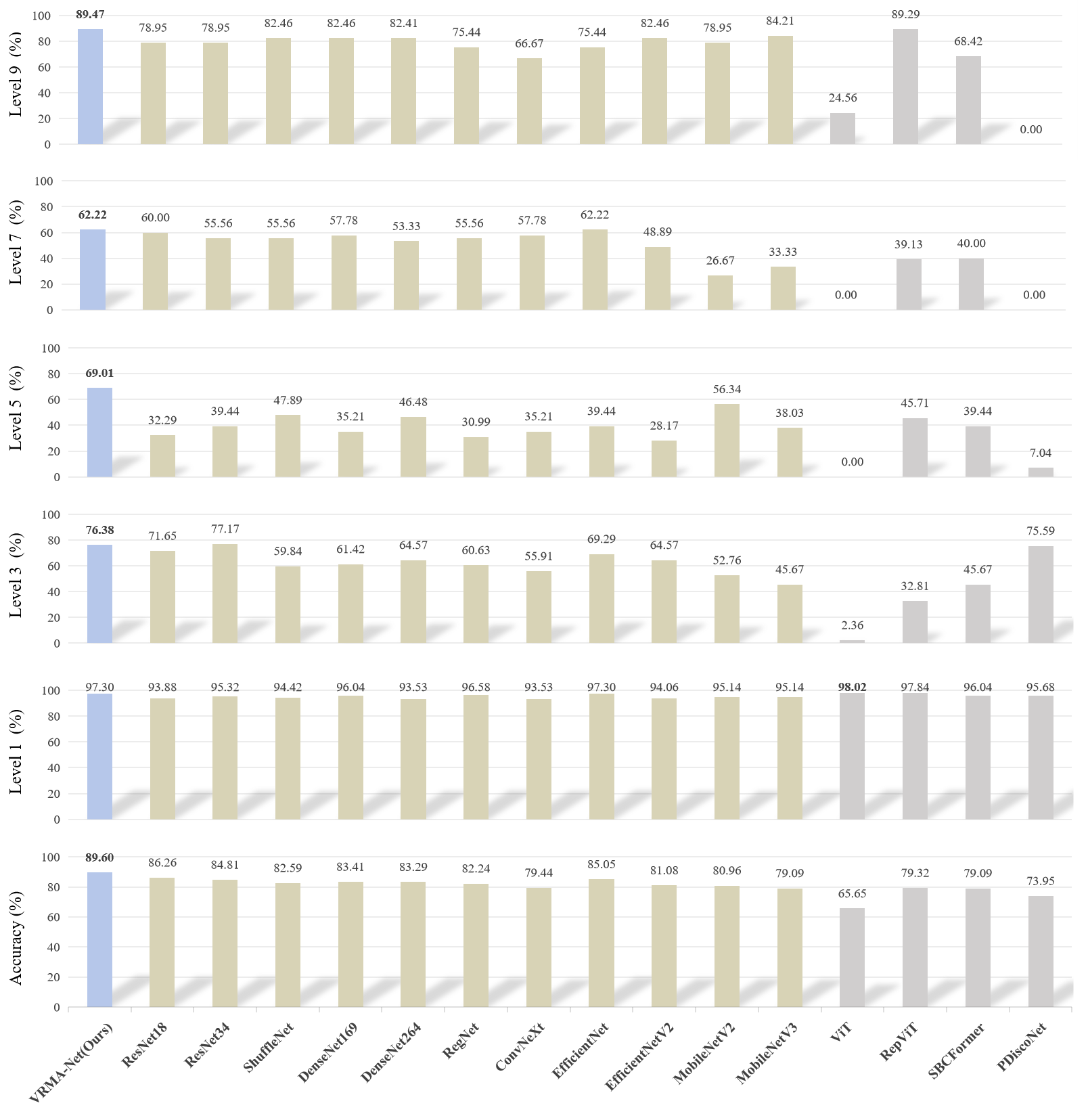
Figure 1: Comparative performance of the proposed FLARE framework versus 15 classification models on the disease severity assessment dataset.
Figure 2 illustrates the visualization of features from different modules. The VAE-Module effectively separates lesion and leaf features, while the VRM-Module models their relationships, suppressing background noise and capturing realistic lesion–leaf distributions. This demonstrates that FLARE achieves accurate and interpretable severity assessment through explicit lesion–leaf awareness and relational modeling.
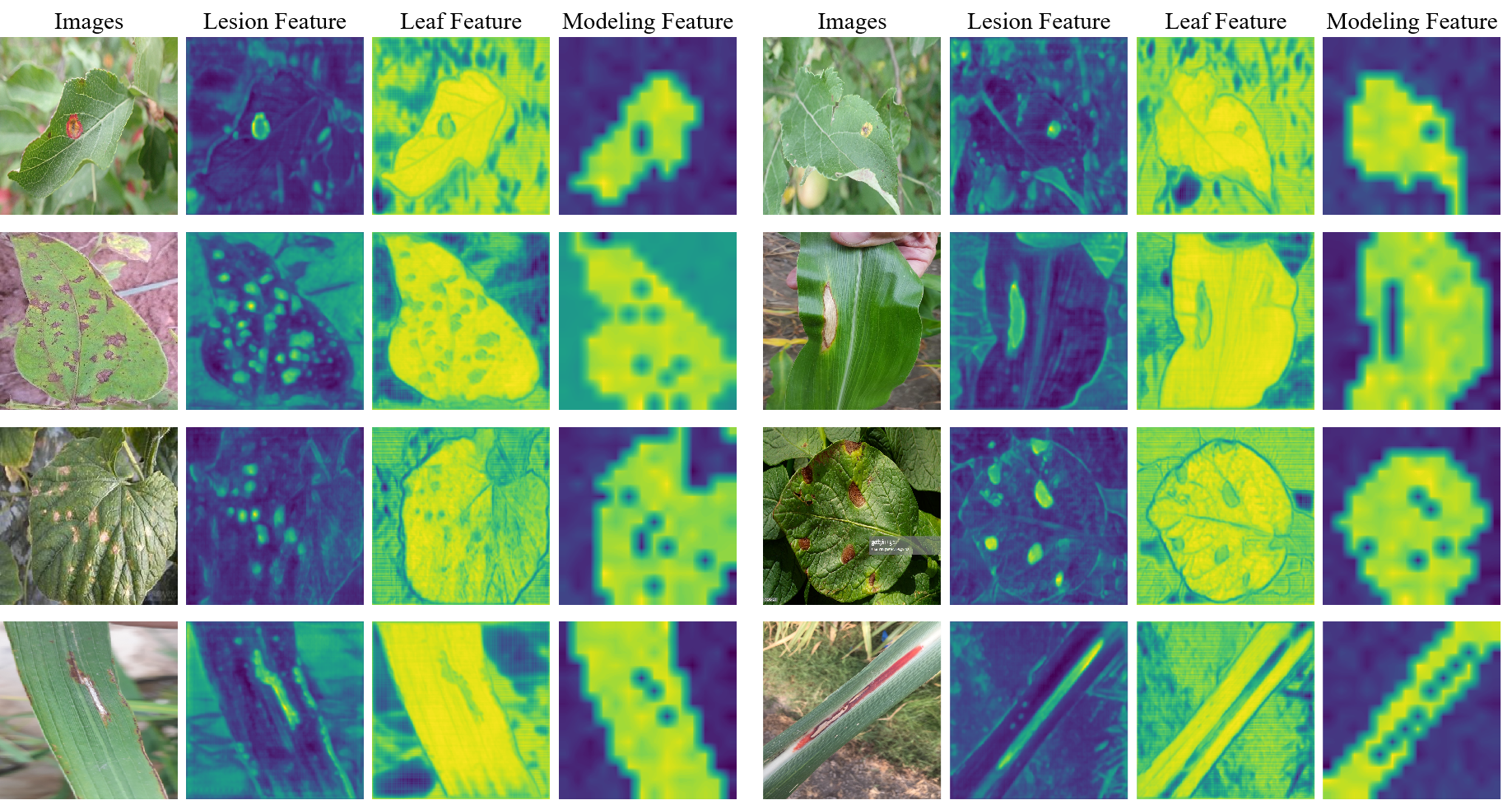

Figure 2: Visualization of features extracted and modeled by key modules in the proposed framework.
Figure 3 presents Grad-CAM visualizations illustrating the model’s attention across network layers. In shallow layers, attention is diffuse, while deeper layers progressively integrate lesion details with the overall leaf structure. This hierarchical attention pattern reflects a human-like perception process, enabling FLARE to achieve more accurate and interpretable severity assessments.
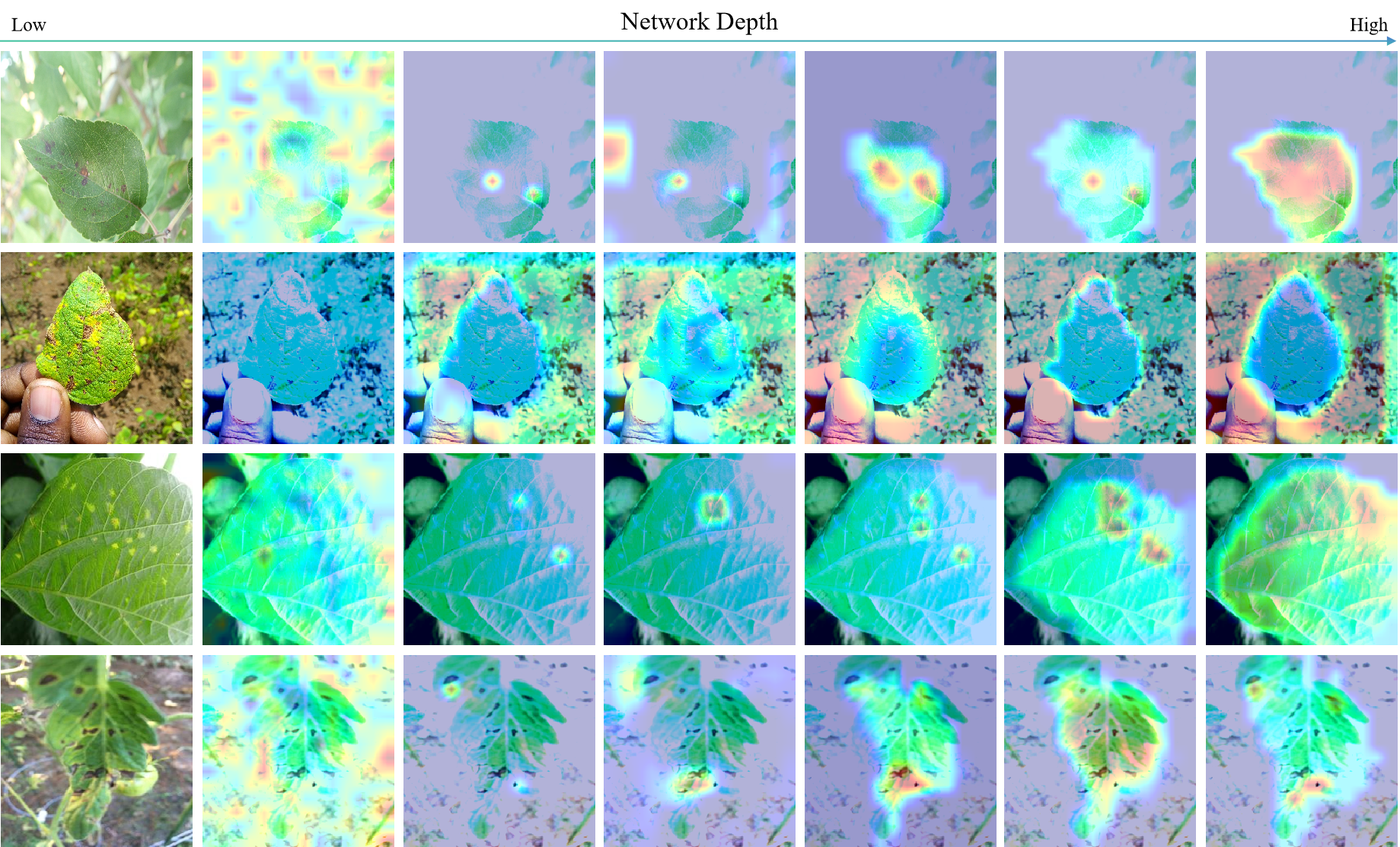

Figure 3: Visualization of model decision-making.
